Large-volume bioimage informatics
Bioimage informatics for large datasets. The imaging methods developed in the group lead to increasing resolution, size and diversity of contrast, which are used to acquire ever larger imaging datasets. This leads to an increasingly active areas of research, bioimage informatics and image based computational biology: how do we handle, process, analyze and visualize Tb sized datasets? How do we use the resulting quantitative measurements to advance biology?
Quantitative geometry. Once large microscopy datasets are fully quantified, the results typically take the form of a large number of geometrical data, point cloud, surfaces, curves or trees that represent cell position and shapes, vesicules trajectories or axonal paths. To answer biological questions, we use datascience/machine learning/AI methods to mine the resulting large database of information. In particular, to ease the building of quantitative biology workflows from those data, we are developing GeNePy3D, a python library to help us solve problems better framed in the context of quantitative geometry (incl. computational geometry and spatial statistics for example).
Image based computational modelling. Beyond bottom-up, data mining, studies, we are increalingly investiguating top down, modelling approaches. To that aim we develop computational models and simulation tailored to specific biological models and questions toward a better integration of data and models. All of those studies are done in deep interdisciplinary collaborations including physicist, data analyst and biologist.
Contact: Anatole Chessel
Related publications
(Preprint) BioImageLoader: Easy Handling of Bioimage Datasets for Machine Learning. Lim, Zhang, Beaurepaire, Chessel, arXiv (2023).
BiaPy: accessible deep learning on bioimages
D. Franco-Barranco, J. A. Andrés-San Román, I. Hidalgo-Cenalmor, L. Backová, A. González-Marfil, C. Caporal, A. Chessel, P. Gómez-Gálvez, L. M. Escudero, D. Wei, A. Muñoz-Barrutia, I. Arganda-Carreras
Nat. Methods (2025).
NU-Net: A Self-Supervised Smart Filter for Enhancing Blobs in Bioimages
S.Lim, E. Beaurepaire, A. Chessel
IEEE ICCV (2023).
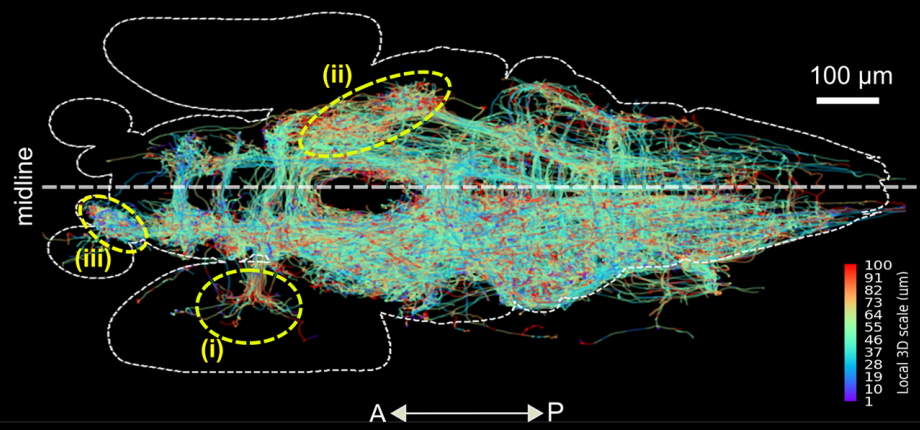
nAdder: A scale-space approach for the 3D analysis of neuronal traces
MS. Phan, K. Matho, E. Beaurepaire, J. Livet, A. Chessel
PLoS Computational Biology (2022).
GeNePy3D: a quantitative geometry python toolbox for bioimaging
MS. Phan, A. Chessel
F1000Research (2020).
Image Data Resource: a bioimage data integration and publication platform
Williams, Moore, Li, Rustici, Tarkowska, Chessel, Leo, Antal, Ferguson, Sarkans, Brazma, Carazo Salas, Swedlow
Nat. Methods (2017).



